
Contributions
The idea of FMODB was born in discussions among Drs. Teruki Honma (RIKEN; the vice chair of the FMO Drug Discovery Consortium; FMODD), Kaori Fukuzawa (Hoshi University; the chair of FMODD), and Shigenori Tanaka (Kobe University; the executive advisor of FMODD) at the launch of FMODD. After the establishment of FMODD, the Honma Laboratory (RIKEN) led the development of FMODB in cooperation with FMODD.
Among the members of the Honma laboratory, Dr. Teruki Honma provided the basic concept and supervised the development of FMODB.
Drs. Chiduru Watanabe and Daisuke Takaya led the actual work on FMODB construction.
Drs. Daisuke Takaya, Chiduru Watanabe, Kikuko Kamisaka, Hirotomo Moriwaki, and Mr. Shunpei Nagase were involved in the development of registration procedures, database schema, and website design.
Drs. Yoshio Okiyama, Chiduru Watanabe, and Kikuko Kamisaka compiled and listed the protein structures related to drug discovery and corresponding binding affinity/activity values.
Drs. Chiduru Watanabe, Kikuko Kamisaka, Hirotomo Moriwaki, and Mr. Shunpei Nagase performed FMO calculations for diverse proteins, including PDB binds data for the initial registration data of FMODB.
Drs. Daisuke Takaya, Hitomi Yuki, and Tomohiro Sato also provided FMO data for on-going drug discovery targets.
Among the members of the FMODD, Dr. Kaori Fukuzawa provided valuable and important opinions regarding all aspects of the construction of FMODB.
Drs. Tomonaga Ozawa (the vice chair of FMODD) and Midori Kamimura (the executive advisor of FMODD) provided useful advice from the point-of-view of the pharmaceutical industry.
Drs. Shigenori Tanaka, Tatsuya Takagi, Norihito Kawashita, Yoichiro Yagi, Noriyuki Kurita, Ken-ichiro Takaba, and Yoshio Okiyama (working group leaders of FMODD) and working group members have registered FMO data obtained from the working group activities as part of the initial data of FMODB.
Many FMODD members provided valuable feedback on the development of FMODB.
Most of the funding used to build and release the initial version of FMODB came from the budgets of the RIKEN Center for Biosystems Dynamics Research and AMED BINDS.
Fragment molecular orbital (FMO) method
It is difficult for conventional ab initio quantum chemical methods to handle biological macromolecules such as proteins and nucleic acids because of the enormous amount of calculation needed. To overcome this difficulty, in the FMO method, macromolecules are divided into small fragments, the electronic states of fragments and fragment pairs are solved in the presence of environmental electrostatic potential from the surroundings, and the entire electronic state is constructed by combining them.
Simultaneously, the interaction energy between fragments (Inter-Fragment Interaction Energy; IFIE or Pair Interaction Energy; PIE) and its energy components (Pair Interaction Energy Decomposition Analysis; PIEDA) can be obtained as by-products of the FMO calculation [1][2][3].
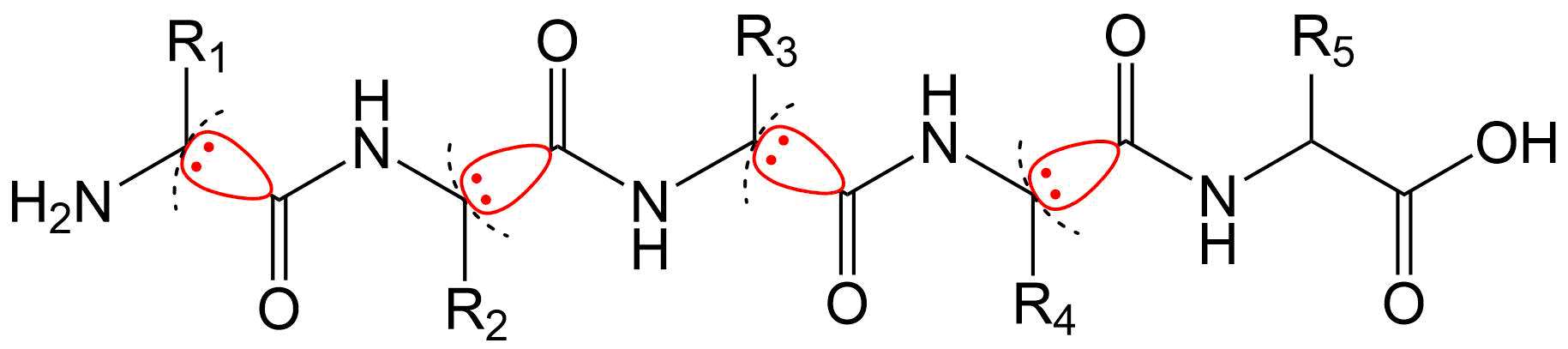
Both IFIE and PIEDA are extensively used in in silico drug design to evaluate the binding (free) energy and molecular interaction between ligands and target proteins. The effects of thermal fluctuation of the molecular structure in solvent can be incorporated by several appropriate methods.
[1] K. Kitaura et al. (1999) Chem. Phys. Lett. 313, 701-709
https://doi.org/10.1016/S0009-2614(99)00937-9
[2] D. G. Fedorov et al. (2012) Phys. Chem. Chem. Phys. 14, 7562-7577.
https://doi.org/10.1039/C2CP23784A
[3] S. Tanaka et al. (2014) Phys. Chem. Chem. Phys. 16, 10310-10344.
https://doi.org/10.1039/C4CP00316K