
FMODB ID: P2LKR
Calculation Name: 4Q1P-B-Xray27
Preferred Name: Galectin-1
Target Type: SINGLE PROTEIN
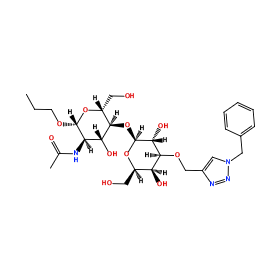
Ligand Name: propyl 2-(acetylamino)-4-o-{3-o-[(1-benzyl-1h-1,2,3-triazol-4-yl)methyl]-beta-d-galactopyranosyl}-2-deoxy-beta-d-glucopyranoside
ligand 3-letter code: 2XT
PDB ID: 4Q1P
Chain ID: B
ChEMBL ID: CHEMBL4915
UniProt ID: P09382
Base Structure: X-ray
Registration Date: 2018-07-26
Reference:
Modeling method
| Optimization | MOE:Amber10:EHT |
|---|---|
| Restraint | OptH |
| Protonation | MOE:Protonate 3D |
| Complement | Missing residues were capped by ACE, NME. Missing atoms were corrected by Structure prepareation by MOE. |
| Water | Apo structure with with a 3 angstrom solvent shell. |
| Procedure | Auto-FMO protocol ver. 1.20180227 |
FMO calculation
| FMO method | FMO2-MP2/6-31G(d) |
|---|---|
| Fragmentation | Auto |
| Number of fragment | 150 |
| LigandCharge | SO4=-2 |
| Software | MIZUHO/ABINIT-MP 3.0 |
Total energy (hartree)
| FMO2-HF: Electronic energy | -1279547.268853 |
|---|---|
| FMO2-HF: Nuclear repulsion | 1222101.211919 |
| FMO2-HF: Total energy | -57446.056934 |
| FMO2-MP2: Total energy | -57606.001661 |
3D Structure
Ligand structure
Ligand Interaction

Ligand binding energy
"| IFIE [kcal/mol] | PIEDA [kcal/mol] | Charge transfer value [e] | |||
|---|---|---|---|---|---|
| IFIE SUMIFIE SUM at MP2 level. | ESElectro static interaction energy. | EXExchange-repulsion energy. | CT+mixCharge transfer and mixing terms energy. | DI(MP2)Dispersion energy. | q(I=>J)Charge transfer value from I to J fragments. |
| N/A | N/A | N/A | N/A | N/A | N/A |
Interactive mode: IFIE and PIEDA for fragment #1(B:1:ACE)
Summations of interaction energy for
fragment #1(B:1:ACE)
| IFIE [kcal/mol] | PIEDA [kcal/mol] | Charge transfer value [e] | |||
|---|---|---|---|---|---|
| IFIE SUMIFIE SUM at MP2 level. | ESElectro static interaction energy. | EXExchange-repulsion energy. | CT+mixCharge transfer and mixing terms energy. | DI(MP2)Dispersion energy. | q(I=>J)Charge transfer value from I to J fragments. |
| -3.172 | -0.423 | 2.182 | -2.8 | -2.131 | -0.021 |
Interaction energy analysis for fragmet #1(B:1:ACE)
| frag_NumFragment number. | ChainChain species. | Res #Residue number. | RES3-letter code of amino acid residue, ligand and solvent molecule. | FCHARGEFormal charge [e]. | q_MullikenFragment charge evaluated by Mulliken charge [e]. | q_NPAFragment charge evaluated by natural population analysis(NPA) charge [e]. | DISTDistance from Ligand [Å]. | TotalIFIE at MP2 level [kcal/mol]. | ESElectro static interaction energy by PIEDA [kcal/mol]. | EXExchange-repulsion energy by PIEDA [kcal/mol]. | CT+mixCharge transfer and mixing terms energy by PIEDA [kcal/mol]. | DI(MP2)Dispersion energy by PIEDA [kcal/mol]. | q(I=>J)Charge transfer value from I to J fragmens [e]. |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 3 | B | 3 | GLY | 0 | 0.044 | 0.035 | 3.776 | 0.829 | 2.073 | -0.015 | -0.607 | -0.622 | -0.001 |
| 4 | B | 4 | LEU | 0 | 0.004 | -0.001 | 5.453 | 0.260 | 0.260 | 0.000 | 0.000 | 0.000 | 0.000 |
| 5 | B | 5 | VAL | 0 | -0.011 | -0.005 | 4.822 | 0.223 | 0.223 | 0.000 | 0.000 | 0.000 | 0.000 |
| 6 | B | 6 | ALA | 0 | -0.021 | -0.004 | 7.652 | -0.117 | -0.117 | 0.000 | 0.000 | 0.000 | 0.000 |
| 7 | B | 7 | SER | 0 | -0.016 | -0.024 | 11.266 | 0.068 | 0.068 | 0.000 | 0.000 | 0.000 | 0.000 |
| 8 | B | 8 | ASN | 0 | -0.033 | -0.016 | 13.374 | -0.025 | -0.025 | 0.000 | 0.000 | 0.000 | 0.000 |
| 9 | B | 9 | LEU | 0 | 0.010 | -0.006 | 13.565 | -0.026 | -0.026 | 0.000 | 0.000 | 0.000 | 0.000 |
| 10 | B | 10 | ASN | 0 | -0.026 | -0.010 | 17.549 | -0.012 | -0.012 | 0.000 | 0.000 | 0.000 | 0.000 |
| 11 | B | 11 | LEU | 0 | 0.032 | 0.048 | 18.594 | -0.014 | -0.014 | 0.000 | 0.000 | 0.000 | 0.000 |
| 12 | B | 12 | LYS | 1 | 0.900 | 0.934 | 20.931 | -0.092 | -0.092 | 0.000 | 0.000 | 0.000 | 0.000 |
| 13 | B | 13 | PRO | 0 | 0.011 | 0.000 | 24.052 | -0.005 | -0.005 | 0.000 | 0.000 | 0.000 | 0.000 |
| 14 | B | 14 | GLY | 0 | -0.039 | -0.009 | 25.565 | -0.007 | -0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 15 | B | 15 | GLU | -1 | -0.815 | -0.893 | 23.681 | 0.076 | 0.076 | 0.000 | 0.000 | 0.000 | 0.000 |
| 16 | B | 16 | CME | 0 | -0.078 | -0.025 | 22.723 | 0.007 | 0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 17 | B | 17 | LEU | 0 | 0.017 | 0.015 | 14.949 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 18 | B | 18 | ARG | 1 | 0.823 | 0.881 | 19.198 | -0.016 | -0.016 | 0.000 | 0.000 | 0.000 | 0.000 |
| 19 | B | 19 | VAL | 0 | 0.042 | 0.017 | 13.174 | -0.006 | -0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 20 | B | 20 | ARG | 1 | 0.945 | 0.985 | 16.583 | 0.085 | 0.085 | 0.000 | 0.000 | 0.000 | 0.000 |
| 21 | B | 21 | GLY | 0 | 0.027 | -0.009 | 14.827 | -0.014 | -0.014 | 0.000 | 0.000 | 0.000 | 0.000 |
| 22 | B | 22 | GLU | -1 | -0.948 | -0.969 | 15.124 | -0.200 | -0.200 | 0.000 | 0.000 | 0.000 | 0.000 |
| 23 | B | 23 | VAL | 0 | 0.008 | 0.014 | 14.585 | -0.051 | -0.051 | 0.000 | 0.000 | 0.000 | 0.000 |
| 24 | B | 24 | ALA | 0 | 0.042 | 0.018 | 14.006 | 0.019 | 0.019 | 0.000 | 0.000 | 0.000 | 0.000 |
| 25 | B | 25 | PRO | 0 | 0.030 | -0.020 | 15.998 | 0.028 | 0.028 | 0.000 | 0.000 | 0.000 | 0.000 |
| 26 | B | 26 | ASP | -1 | -0.957 | -0.964 | 16.204 | -0.305 | -0.305 | 0.000 | 0.000 | 0.000 | 0.000 |
| 27 | B | 27 | ALA | 0 | -0.034 | -0.005 | 14.943 | -0.007 | -0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 28 | B | 28 | LYS | 1 | 0.912 | 0.951 | 13.930 | 0.260 | 0.260 | 0.000 | 0.000 | 0.000 | 0.000 |
| 29 | B | 29 | SER | 0 | -0.010 | -0.022 | 10.531 | 0.089 | 0.089 | 0.000 | 0.000 | 0.000 | 0.000 |
| 30 | B | 30 | PHE | 0 | -0.005 | 0.023 | 9.496 | -0.090 | -0.090 | 0.000 | 0.000 | 0.000 | 0.000 |
| 31 | B | 31 | VAL | 0 | 0.011 | 0.011 | 6.693 | 0.116 | 0.116 | 0.000 | 0.000 | 0.000 | 0.000 |
| 32 | B | 32 | LEU | 0 | -0.010 | 0.004 | 7.986 | -0.072 | -0.072 | 0.000 | 0.000 | 0.000 | 0.000 |
| 33 | B | 33 | ASN | 0 | 0.023 | 0.021 | 6.147 | -0.209 | -0.209 | 0.000 | 0.000 | 0.000 | 0.000 |
| 34 | B | 34 | LEU | 0 | 0.045 | 0.013 | 9.445 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 35 | B | 35 | GLY | 0 | 0.049 | 0.006 | 12.472 | 0.007 | 0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 36 | B | 36 | LYS | 1 | 0.970 | 0.999 | 14.361 | -0.322 | -0.322 | 0.000 | 0.000 | 0.000 | 0.000 |
| 37 | B | 37 | ASP | -1 | -0.847 | -0.934 | 11.991 | 0.476 | 0.476 | 0.000 | 0.000 | 0.000 | 0.000 |
| 38 | B | 38 | SER | 0 | -0.001 | -0.001 | 7.565 | -0.093 | -0.093 | 0.000 | 0.000 | 0.000 | 0.000 |
| 39 | B | 39 | ASN | 0 | -0.038 | -0.024 | 9.427 | 0.090 | 0.090 | 0.000 | 0.000 | 0.000 | 0.000 |
| 40 | B | 40 | ASN | 0 | -0.045 | -0.013 | 12.292 | -0.117 | -0.117 | 0.000 | 0.000 | 0.000 | 0.000 |
| 41 | B | 41 | LEU | 0 | -0.036 | -0.011 | 9.162 | 0.003 | 0.003 | 0.000 | 0.000 | 0.000 | 0.000 |
| 42 | B | 42 | CYS | 0 | 0.027 | 0.015 | 13.557 | -0.031 | -0.031 | 0.000 | 0.000 | 0.000 | 0.000 |
| 43 | B | 43 | LEU | 0 | 0.012 | -0.002 | 14.213 | -0.037 | -0.037 | 0.000 | 0.000 | 0.000 | 0.000 |
| 44 | B | 44 | HIS | 0 | -0.071 | -0.023 | 8.622 | -0.003 | -0.003 | 0.000 | 0.000 | 0.000 | 0.000 |
| 45 | B | 45 | PHE | 0 | 0.052 | 0.004 | 11.583 | -0.050 | -0.050 | 0.000 | 0.000 | 0.000 | 0.000 |
| 46 | B | 46 | ASN | 0 | 0.012 | 0.000 | 11.411 | 0.013 | 0.013 | 0.000 | 0.000 | 0.000 | 0.000 |
| 47 | B | 47 | PRO | 0 | 0.016 | 0.018 | 12.634 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 48 | B | 48 | ARG | 1 | 0.813 | 0.899 | 13.258 | -0.098 | -0.098 | 0.000 | 0.000 | 0.000 | 0.000 |
| 49 | B | 49 | PHE | 0 | 0.030 | 0.016 | 15.174 | 0.015 | 0.015 | 0.000 | 0.000 | 0.000 | 0.000 |
| 50 | B | 50 | ASN | 0 | -0.062 | -0.045 | 17.137 | 0.014 | 0.014 | 0.000 | 0.000 | 0.000 | 0.000 |
| 51 | B | 51 | ALA | 0 | 0.027 | 0.012 | 14.240 | -0.026 | -0.026 | 0.000 | 0.000 | 0.000 | 0.000 |
| 52 | B | 52 | HIS | 0 | -0.008 | -0.005 | 11.466 | 0.033 | 0.033 | 0.000 | 0.000 | 0.000 | 0.000 |
| 53 | B | 53 | GLY | 0 | 0.006 | 0.009 | 16.935 | 0.017 | 0.017 | 0.000 | 0.000 | 0.000 | 0.000 |
| 54 | B | 54 | ASP | -1 | -0.790 | -0.857 | 18.313 | 0.033 | 0.033 | 0.000 | 0.000 | 0.000 | 0.000 |
| 55 | B | 55 | ALA | 0 | 0.014 | -0.013 | 19.571 | -0.017 | -0.017 | 0.000 | 0.000 | 0.000 | 0.000 |
| 56 | B | 56 | ASN | 0 | -0.045 | -0.019 | 20.639 | 0.002 | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 57 | B | 57 | THR | 0 | -0.003 | -0.003 | 19.557 | 0.011 | 0.011 | 0.000 | 0.000 | 0.000 | 0.000 |
| 58 | B | 58 | ILE | 0 | 0.025 | 0.010 | 16.560 | -0.007 | -0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 59 | B | 59 | VAL | 0 | -0.026 | 0.003 | 15.311 | 0.008 | 0.008 | 0.000 | 0.000 | 0.000 | 0.000 |
| 60 | B | 60 | CYS | 0 | -0.003 | 0.002 | 15.042 | 0.015 | 0.015 | 0.000 | 0.000 | 0.000 | 0.000 |
| 61 | B | 61 | ASN | 0 | 0.022 | -0.006 | 14.399 | -0.013 | -0.013 | 0.000 | 0.000 | 0.000 | 0.000 |
| 62 | B | 62 | SER | 0 | 0.042 | 0.025 | 15.078 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 63 | B | 63 | LYS | 1 | 0.872 | 0.944 | 9.467 | -0.813 | -0.813 | 0.000 | 0.000 | 0.000 | 0.000 |
| 64 | B | 64 | ASP | -1 | -0.870 | -0.929 | 14.801 | 0.262 | 0.262 | 0.000 | 0.000 | 0.000 | 0.000 |
| 65 | B | 65 | GLY | 0 | 0.057 | 0.027 | 17.072 | -0.011 | -0.011 | 0.000 | 0.000 | 0.000 | 0.000 |
| 66 | B | 66 | GLY | 0 | -0.057 | -0.035 | 13.582 | -0.011 | -0.011 | 0.000 | 0.000 | 0.000 | 0.000 |
| 67 | B | 67 | ALA | 0 | -0.036 | -0.011 | 14.318 | 0.018 | 0.018 | 0.000 | 0.000 | 0.000 | 0.000 |
| 68 | B | 68 | TRP | 0 | -0.032 | -0.028 | 9.735 | 0.028 | 0.028 | 0.000 | 0.000 | 0.000 | 0.000 |
| 69 | B | 69 | GLY | 0 | 0.023 | 0.025 | 15.975 | -0.031 | -0.031 | 0.000 | 0.000 | 0.000 | 0.000 |
| 70 | B | 70 | THR | 0 | 0.001 | -0.003 | 18.545 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 71 | B | 71 | GLU | -1 | -0.870 | -0.924 | 16.652 | 0.130 | 0.130 | 0.000 | 0.000 | 0.000 | 0.000 |
| 72 | B | 72 | GLN | 0 | -0.105 | -0.072 | 19.179 | -0.008 | -0.008 | 0.000 | 0.000 | 0.000 | 0.000 |
| 73 | B | 73 | ARG | 1 | 0.762 | 0.856 | 18.906 | -0.058 | -0.058 | 0.000 | 0.000 | 0.000 | 0.000 |
| 74 | B | 74 | GLU | -1 | -0.816 | -0.887 | 20.251 | 0.069 | 0.069 | 0.000 | 0.000 | 0.000 | 0.000 |
| 75 | B | 75 | ALA | 0 | -0.024 | -0.014 | 22.794 | -0.006 | -0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 76 | B | 76 | VAL | 0 | -0.029 | -0.010 | 23.315 | -0.006 | -0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 77 | B | 77 | PHE | 0 | 0.006 | -0.015 | 18.011 | 0.004 | 0.004 | 0.000 | 0.000 | 0.000 | 0.000 |
| 78 | B | 78 | PRO | 0 | -0.002 | 0.023 | 21.445 | -0.007 | -0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 79 | B | 79 | PHE | 0 | 0.015 | -0.001 | 17.785 | -0.005 | -0.005 | 0.000 | 0.000 | 0.000 | 0.000 |
| 80 | B | 80 | GLN | 0 | 0.010 | -0.008 | 20.705 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 81 | B | 81 | PRO | 0 | 0.003 | 0.010 | 18.777 | -0.008 | -0.008 | 0.000 | 0.000 | 0.000 | 0.000 |
| 82 | B | 82 | GLY | 0 | -0.020 | 0.004 | 18.848 | 0.014 | 0.014 | 0.000 | 0.000 | 0.000 | 0.000 |
| 83 | B | 83 | SER | 0 | -0.042 | -0.014 | 19.596 | 0.019 | 0.019 | 0.000 | 0.000 | 0.000 | 0.000 |
| 84 | B | 84 | VAL | 0 | -0.026 | -0.017 | 19.113 | -0.018 | -0.018 | 0.000 | 0.000 | 0.000 | 0.000 |
| 85 | B | 85 | ALA | 0 | 0.011 | 0.006 | 17.742 | 0.014 | 0.014 | 0.000 | 0.000 | 0.000 | 0.000 |
| 86 | B | 86 | GLU | -1 | -0.802 | -0.915 | 18.070 | -0.033 | -0.033 | 0.000 | 0.000 | 0.000 | 0.000 |
| 87 | B | 87 | VAL | 0 | -0.018 | 0.011 | 16.095 | 0.005 | 0.005 | 0.000 | 0.000 | 0.000 | 0.000 |
| 88 | B | 88 | CME | 0 | -0.103 | -0.059 | 18.403 | 0.001 | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 89 | B | 89 | ILE | 0 | 0.003 | 0.003 | 15.847 | 0.005 | 0.005 | 0.000 | 0.000 | 0.000 | 0.000 |
| 90 | B | 90 | THR | 0 | 0.039 | 0.021 | 19.777 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 91 | B | 91 | PHE | 0 | -0.048 | -0.029 | 20.985 | 0.009 | 0.009 | 0.000 | 0.000 | 0.000 | 0.000 |
| 92 | B | 92 | ASP | -1 | -0.815 | -0.902 | 23.099 | 0.077 | 0.077 | 0.000 | 0.000 | 0.000 | 0.000 |
| 93 | B | 93 | GLN | 0 | 0.018 | 0.004 | 25.015 | 0.011 | 0.011 | 0.000 | 0.000 | 0.000 | 0.000 |
| 94 | B | 94 | ALA | 0 | -0.018 | -0.004 | 26.626 | 0.006 | 0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 95 | B | 95 | ASN | 0 | -0.052 | -0.033 | 24.074 | 0.008 | 0.008 | 0.000 | 0.000 | 0.000 | 0.000 |
| 96 | B | 96 | LEU | 0 | 0.037 | 0.023 | 17.737 | 0.001 | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 97 | B | 97 | THR | 0 | -0.055 | -0.036 | 21.789 | -0.010 | -0.010 | 0.000 | 0.000 | 0.000 | 0.000 |
| 98 | B | 98 | VAL | 0 | 0.009 | 0.003 | 18.190 | 0.003 | 0.003 | 0.000 | 0.000 | 0.000 | 0.000 |
| 99 | B | 99 | LYS | 1 | 0.901 | 0.952 | 20.871 | -0.035 | -0.035 | 0.000 | 0.000 | 0.000 | 0.000 |
| 100 | B | 100 | LEU | 0 | -0.024 | -0.022 | 19.178 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 101 | B | 101 | PRO | 0 | 0.051 | 0.023 | 20.469 | 0.002 | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 102 | B | 102 | ASP | -1 | -0.912 | -0.965 | 23.642 | -0.031 | -0.031 | 0.000 | 0.000 | 0.000 | 0.000 |
| 103 | B | 103 | GLY | 0 | -0.041 | -0.015 | 26.838 | 0.002 | 0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 104 | B | 104 | TYR | 0 | -0.021 | 0.005 | 24.708 | 0.006 | 0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 105 | B | 105 | GLU | -1 | -0.927 | -0.982 | 23.996 | 0.035 | 0.035 | 0.000 | 0.000 | 0.000 | 0.000 |
| 106 | B | 106 | PHE | 0 | -0.021 | 0.000 | 18.344 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 107 | B | 107 | LYS | 1 | 0.938 | 0.963 | 23.248 | -0.054 | -0.054 | 0.000 | 0.000 | 0.000 | 0.000 |
| 108 | B | 108 | PHE | 0 | 0.020 | 0.019 | 17.889 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 109 | B | 109 | PRO | 0 | 0.039 | 0.023 | 21.958 | 0.004 | 0.004 | 0.000 | 0.000 | 0.000 | 0.000 |
| 110 | B | 110 | ASN | 0 | -0.036 | -0.021 | 21.313 | 0.036 | 0.036 | 0.000 | 0.000 | 0.000 | 0.000 |
| 111 | B | 111 | ARG | 1 | 0.841 | 0.896 | 19.241 | -0.173 | -0.173 | 0.000 | 0.000 | 0.000 | 0.000 |
| 112 | B | 112 | LEU | 0 | -0.009 | 0.000 | 16.832 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 113 | B | 113 | ASN | 0 | -0.044 | -0.010 | 21.489 | -0.011 | -0.011 | 0.000 | 0.000 | 0.000 | 0.000 |
| 114 | B | 114 | LEU | 0 | -0.049 | -0.019 | 17.321 | 0.001 | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 115 | B | 115 | GLU | -1 | -0.879 | -0.947 | 22.027 | 0.102 | 0.102 | 0.000 | 0.000 | 0.000 | 0.000 |
| 116 | B | 116 | ALA | 0 | -0.045 | -0.040 | 21.452 | -0.013 | -0.013 | 0.000 | 0.000 | 0.000 | 0.000 |
| 117 | B | 117 | ILE | 0 | -0.026 | 0.008 | 15.536 | 0.027 | 0.027 | 0.000 | 0.000 | 0.000 | 0.000 |
| 118 | B | 118 | ASN | 0 | -0.026 | -0.029 | 16.439 | -0.035 | -0.035 | 0.000 | 0.000 | 0.000 | 0.000 |
| 119 | B | 119 | TYR | 0 | 0.006 | 0.014 | 6.195 | 0.067 | 0.067 | 0.000 | 0.000 | 0.000 | 0.000 |
| 120 | B | 120 | MET | 0 | 0.029 | 0.007 | 10.110 | -0.043 | -0.043 | 0.000 | 0.000 | 0.000 | 0.000 |
| 121 | B | 121 | ALA | 0 | -0.024 | 0.003 | 5.006 | 0.048 | 0.048 | 0.000 | 0.000 | 0.000 | 0.000 |
| 122 | B | 122 | ALA | 0 | -0.001 | 0.006 | 6.150 | 0.006 | 0.006 | 0.000 | 0.000 | 0.000 | 0.000 |
| 123 | B | 123 | ASP | -1 | -0.852 | -0.936 | 2.543 | -4.209 | -3.086 | 2.146 | -1.967 | -1.303 | -0.019 |
| 124 | B | 124 | GLY | 0 | -0.027 | -0.007 | 4.905 | 0.466 | 0.505 | -0.001 | -0.003 | -0.034 | 0.000 |
| 125 | B | 125 | ASP | -1 | -0.771 | -0.884 | 7.722 | -0.530 | -0.530 | 0.000 | 0.000 | 0.000 | 0.000 |
| 126 | B | 126 | PHE | 0 | -0.026 | -0.028 | 10.018 | 0.122 | 0.122 | 0.000 | 0.000 | 0.000 | 0.000 |
| 127 | B | 127 | LYS | 1 | 0.943 | 0.989 | 10.116 | 0.248 | 0.248 | 0.000 | 0.000 | 0.000 | 0.000 |
| 128 | B | 128 | ILE | 0 | 0.032 | 0.021 | 10.175 | 0.071 | 0.071 | 0.000 | 0.000 | 0.000 | 0.000 |
| 129 | B | 129 | LYS | 1 | 0.871 | 0.928 | 12.987 | 0.186 | 0.186 | 0.000 | 0.000 | 0.000 | 0.000 |
| 130 | B | 130 | CME | 0 | -0.170 | -0.099 | 16.518 | 0.015 | 0.015 | 0.000 | 0.000 | 0.000 | 0.000 |
| 131 | B | 131 | VAL | 0 | 0.031 | 0.019 | 14.066 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 132 | B | 132 | ALA | 0 | 0.000 | -0.002 | 17.348 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 133 | B | 133 | PHE | 0 | -0.033 | -0.019 | 17.203 | 0.010 | 0.010 | 0.000 | 0.000 | 0.000 | 0.000 |
| 134 | B | 134 | ASP | -2 | -1.647 | -1.803 | 22.166 | 0.065 | 0.065 | 0.000 | 0.000 | 0.000 | 0.000 |
| 135 | B | 201 | SO4 | -2 | -1.945 | -1.970 | 14.358 | -0.846 | -0.846 | 0.000 | 0.000 | 0.000 | 0.000 |
| 136 | B | 202 | 2XT | 0 | 0.001 | 0.037 | 26.349 | -0.007 | -0.007 | 0.000 | 0.000 | 0.000 | 0.000 |
| 137 | A | 323 | HOH | 0 | -0.023 | -0.012 | 3.475 | 0.728 | 1.071 | 0.052 | -0.223 | -0.172 | -0.001 |
| 138 | A | 339 | HOH | 0 | -0.026 | -0.013 | 20.454 | -0.001 | -0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 139 | A | 340 | HOH | 0 | -0.028 | -0.013 | 22.988 | 0.001 | 0.001 | 0.000 | 0.000 | 0.000 | 0.000 |
| 140 | A | 352 | HOH | 0 | -0.054 | -0.032 | 23.681 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 141 | A | 356 | HOH | 0 | 0.013 | 0.009 | 18.023 | -0.004 | -0.004 | 0.000 | 0.000 | 0.000 | 0.000 |
| 142 | A | 387 | HOH | 0 | -0.054 | -0.042 | 11.890 | 0.010 | 0.010 | 0.000 | 0.000 | 0.000 | 0.000 |
| 143 | A | 393 | HOH | 0 | 0.009 | 0.000 | 14.873 | 0.008 | 0.008 | 0.000 | 0.000 | 0.000 | 0.000 |
| 144 | B | 347 | HOH | 0 | -0.055 | -0.030 | 15.106 | 0.016 | 0.016 | 0.000 | 0.000 | 0.000 | 0.000 |
| 145 | B | 370 | HOH | 0 | 0.018 | 0.023 | 9.772 | 0.047 | 0.047 | 0.000 | 0.000 | 0.000 | 0.000 |
| 146 | B | 374 | HOH | 0 | -0.024 | -0.052 | 23.973 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 147 | B | 377 | HOH | 0 | 0.024 | 0.015 | 31.955 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 | 0.000 |
| 148 | B | 379 | HOH | 0 | 0.008 | 0.002 | 20.026 | -0.002 | -0.002 | 0.000 | 0.000 | 0.000 | 0.000 |
| 149 | B | 382 | HOH | 0 | -0.008 | -0.006 | 7.681 | 0.141 | 0.141 | 0.000 | 0.000 | 0.000 | 0.000 |
| 150 | B | 415 | HOH | 0 | -0.011 | -0.003 | 25.042 | -0.003 | -0.003 | 0.000 | 0.000 | 0.000 | 0.000 |